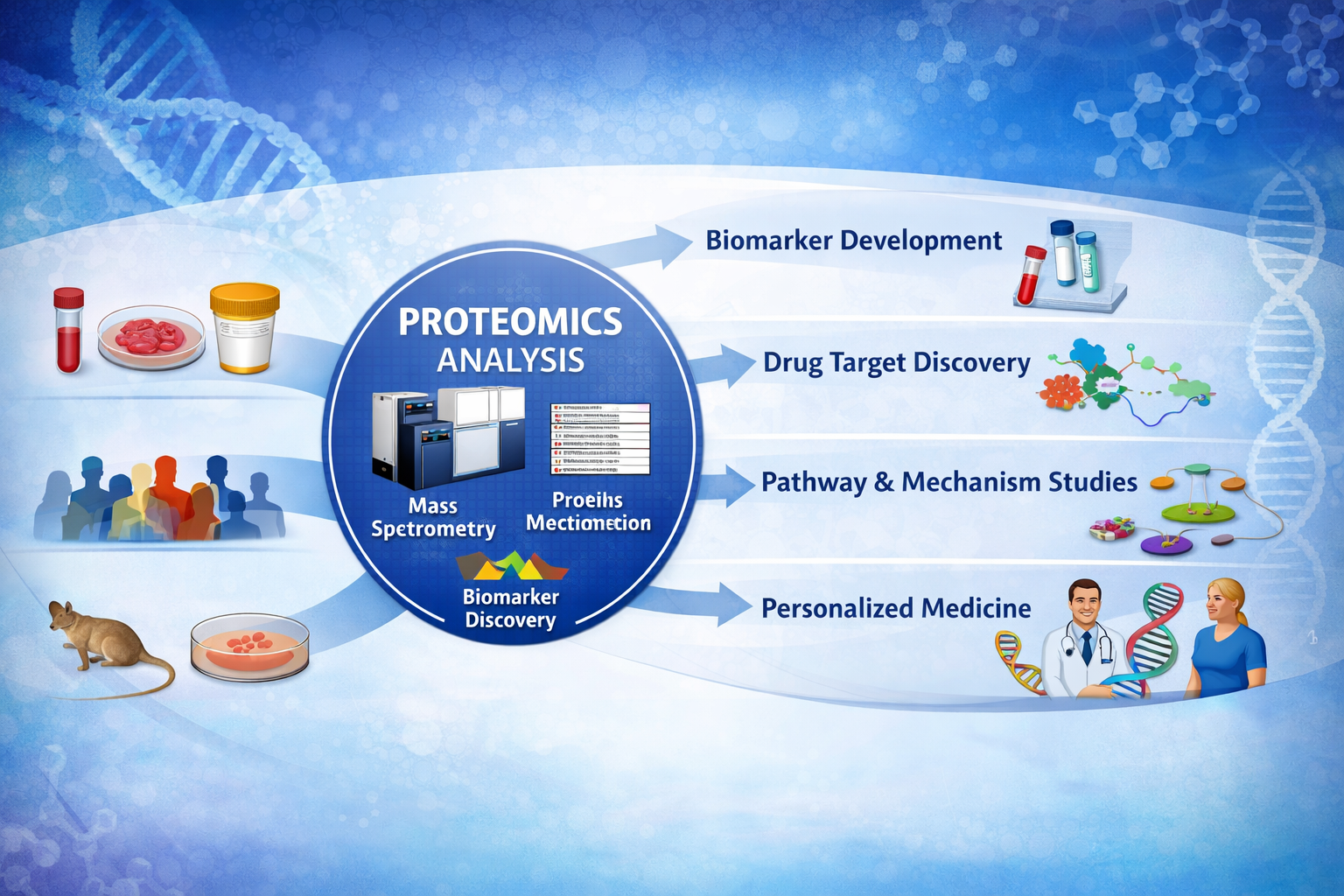
Proteomics has matured significantly as a research tool, and its integration into translational research programs is now a practical and strategically important consideration for organisations seeking deeper biological insight. While genomic and transcriptomic data reveal what is encoded or expressed, protein-level measurements reflect the functional output of biological systems—making proteomics services an essential complement to molecular profiling strategies focused on biomarker discovery, target identification, and disease mechanism characterisation.
The translational value of protein-level data
Gene expression and protein abundance do not always correlate directly. Post-translational modifications, protein stability, and context-dependent regulation mean that understanding biology at the protein level often reveals mechanistic information that transcriptomics alone cannot capture. In translational research, this distinction matters: protein biomarkers are frequently the most actionable indicators of disease state, treatment response, or patient stratification relevant to clinical decision-making.
Selecting the right proteomics approach
Proteomics services encompass a range of technologies with different performance characteristics. Affinity-based multiplex platforms—such as proximity extension assay technologies—enable simultaneous quantification of hundreds to thousands of proteins from small sample volumes with high sensitivity and specificity. Mass spectrometry-based approaches offer broad, unbiased coverage and are well-suited for discovery applications. Selecting the appropriate platform depends on study objectives, sample type and volume, required throughput, and the degree of prior knowledge available to guide panel selection.
Study design considerations for proteomics integration
Successful integration of proteomics into translational programs requires careful study design. Sample collection, handling, and storage conditions have a significant impact on protein stability and measurement quality—standardised biobanking protocols are essential. Statistical power calculations should account for the multiplicity of protein targets being evaluated and the expected effect sizes in the biological system under investigation. Pre-specifying primary analytes and hypotheses also reduces the risk of spurious findings from large-scale protein profiling datasets.
Data integration across omics layers
Proteomics data becomes most powerful when integrated with complementary genomic, transcriptomic, or clinical datasets. Multi-omics integration allows researchers to identify convergent signals across biological layers and build more robust mechanistic models. Achieving meaningful integration requires harmonised metadata, consistent sample provenance, and analytical frameworks capable of handling high-dimensional data from heterogeneous sources. Building these capabilities early in program design reduces friction when integrating data downstream.
Regulatory and reporting considerations
For translational programs with a regulatory trajectory, proteomics data generation and analysis must meet appropriate standards for documentation, reproducibility, and analytical validation. Partnering with proteomics services providers experienced in regulated research environments helps ensure that study data will support future regulatory submissions and withstand scientific scrutiny.
Conclusion
Proteomics services offer translational research programs a powerful window into the functional biology underlying disease and treatment response. By integrating protein-level profiling thoughtfully—with appropriate platform selection, rigorous study design, and multi-omics integration strategies—research teams can generate insights that accelerate the path from biological discovery to clinical application.